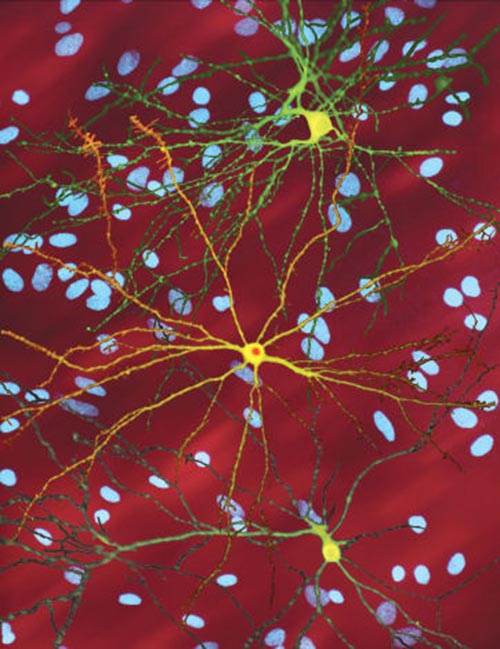
A montage of three images of single striatal neurons transfected with a disease-associated version of huntingtin, the protein that causes Huntington’s disease. Nuclei of untransfected neurons are seen in the background (blue). The neuron in the center (yellow) contains an abnormal intracellular accumulation of huntingtin called an inclusion body (orange). Photo: Dr. Steven Finkbeiner, Gladstone Institute of Neurological Disease, The Taube-Koret Center for Huntington’s Disease Research, and the University of California San Francisco
Patients with Huntington’s disease, a fatal genetic illness that causes the breakdown of nerve cells in the brain, have up to 80 percent less cancer than the general population.
Northwestern Medicine scientists have discovered why Huntington’s is so toxic to cancer cells and harnessed it for a novel approach to treat cancer, a new study reports.
Huntington’s is caused by an over abundance of a certain type of repeating RNA sequences in one gene, huntingtin, present in every cell. The defect that causes the disease also is highly toxic to tumor cells. These repeating sequences — in the form of so-called small interfering RNAs — attack genes in the cell that are critical for survival. Nerve cells in the brain are vulnerable to this form of cell death, however, cancer cells appear to be much more susceptible.
“This molecule is a super assassin against all tumor cells,” said senior author Marcus Peter, the Tom D. Spies Professor of Cancer Metabolism at Northwestern University Feinberg School of Medicine. “We’ve never seen anything this powerful.”
Huntington’s disease deteriorates a person’s physical and mental abilities during their prime working years and has no cure.
The study will be published Feb. 12 in the journal EMBO Reports.
To test the super assassin molecule in a treatment situation, Peter collaborated with Dr. Shad Thaxton, associate professor of urology at Feinberg, to deliver the molecule in nanoparticles to mice with human ovarian cancer. The treatment significantly reduced the tumor growth with no toxicity to the mice, Peter said. Importantly, the tumors did not develop resistance to this form of cancer treatment.
Peter and Thaxton are now refining the delivery method to increase its efficacy in reaching the tumor. The other challenge for the scientists is figuring out how to stabilize the nanoparticles, so they can be stored.
First and co-corresponding author Andrea Murmann, research assistant professor in medicine at Feinberg, also used the molecule to treat human and mouse ovarian, breast, prostate, liver, brain, lung, skin and colon cancer cell lines. The molecule killed all cancer cells in both species.
The Huntington’s cancer weapon was discovered by Murmann, who had worked with Peter on earlier research that identified an ancient kill-switch present in all cells that destroys cancer.
“I thought maybe there is a situation where this kill switch is overactive in certain people, and where it could cause loss of tissues,” Murmann said. “These patients would not only have a disease with an RNA component, but they also had to have less cancer.”
She started searching for diseases that have a lower rate of cancer and had a suspected contribution of RNA to disease pathology. Huntington’s was the most prominent.
When she looked at the repeating sequences in huntingtin, the gene that causes the disease, she saw a similar composition to the earlier kill switch Peter had found. Both were rich in the C and G nucleotides (molecules that form the building blocks of DNA and RNA).
“Toxicity goes together with C and G richness,” Murmann said. “Those similarities triggered our curiosity.”
In the case of people who have Huntington’s, the gene huntingtin has too many repeating sequences of the triplet sequence CAG. The longer the repeating sequence, the earlier they will develop the disease.
“We believe a short-term treatment cancer therapy for a few weeks might be possible, where we could treat a patient to kill the cancer cells without causing the neurological issues that Huntington’s patients suffer from,” Peter said.
Peter also is co-leader of the Translational Research in Solid Tumors Program at the Robert H. Lurie Comprehensive Cancer Center of Northwestern University.
Huntington’s patients have a lifetime exposure to these toxic RNA sequences, but generally don’t develop symptoms of the disease until age 40, he noted.
Every child of a parent with Huntington’s has 50/50 chance of carrying the faulty gene. Today, there are approximately 30,000 symptomatic Americans and more than 200,000 at-risk of inheriting the disease.
Filed Under: Genomics/Proteomics




