
[Image courtesy of DeepMind]
“When we launched the database last July, it was sort of recognized as a pretty big leap forward for biology,” said Demis Hassabis, founder and CEO of DeepMind, in a press conference. The release last year included roughly 350,000 high-quality predictions, including all proteins in the human body.
The continued advances of AlphaFold make it possible to search for the 3-D structure of proteins “almost as easily as doing a keyword Google search,” Hassabis said.
AlphaFold has made the data freely available to users for any purpose.
Here are seven examples of how the AI system is helping fuel progress in life sciences research.
1. Accelerating discovery of drugs for neglected diseases
The Drugs for Neglected Diseases Initiative (DNDi) is using AlphaFold to create new medicines for neglected diseases that disproportionately affect those in developing countries. Specifically, the protein-folding database could accelerate the identification of molecules to treat neglected diseases.
Examples of such diseases include Chagas disease, which is spread by the parasite Trypanosoma cruzi. Chagas disease is common in Mexico, Central America and South America.
Another example is leishmaniasis, a parasitic disease found in tropical and subtropical regions as well as southern Europe.
DNDi now has more than 20 new chemical entities in its portfolio.
DNDi is also working to enable researchers in low-income countries to play a more active role in drug discovery.
2. Combating antibiotic resistance
In 2019, CDC estimated that antimicrobial resistance costs the U.S. economy $55 billion annually, including $20 billion in healthcare costs and another $35 billion in lost productivity. Antibiotic resistance is also fueling the emergence of ‘superbugs.’
Professors Marcelo Sousa and Megan Mitchell from the University of Colorado, Boulder, are using AlphaFold to study proteins involved in antibiotic resistance. AlphaFold has helped the researchers identify protein structures they could confirm with crystallography. For example, Sousa used this technology to identify a bacterial protein structure in approximately 30 minutes that had proven hard to identify for 10 years.
3. Studying the nuclear pore complex
An international research team is working to uncover the secrets of nuclear pore complexes (NPCs), which could be the largest complexes in human cells. NPCs are involved in the transport between the nucleus and cytoplasm and various cellular processes.
Using AlphaFold with Cryo-EM, the researchers have built an almost complete structure of the NPC scaffold. The research project could pave the way for an improved understanding of NPC function, biogenesis and regulation.
Researchers involved in the project hailed from the Max Planck Institute of Biophysics; European Molecular Biology Laboratory; Harvard Medical School; University of California, Berkeley; SLAC National Accelerator Laboratory; Westlake University; and Tsinghua University.
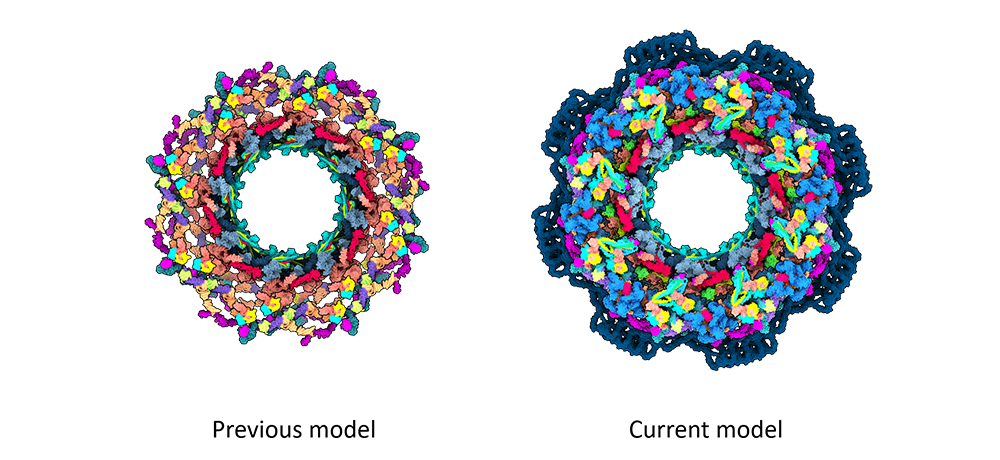
[Image courtesy of DeepMind]
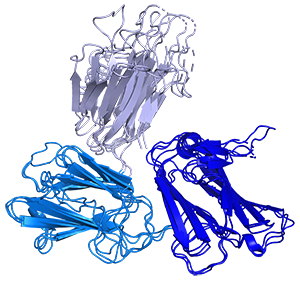
4. Developing a novel malaria vaccine
[Image courtesy of DeepMind]
5. Shedding light on genetic variation
Pedro Beltrao, a professor at ETH Zurich, is working to uncover how DNA changes inform traits, including predisposition to some diseases. “Ultimately, what we’d like to have eventually is a model that tells us exactly how a person is going to change, or what traits they will have if they carry a mutation in a particular position in their DNA,” Beltrao said in a statement.
Beltrao used AlphaFold to study proteins across evolutionary history, helping uncover the proteins present in the ancestral cell. Deciphering which DNA mutations are involved in traits also involves determining which mutations have essentially no impact on proteins. The research also involved studying how multiple proteins work together to perform functions.
“We can now trace back the evolution of proteins for longer periods of the evolutionary timescale,” Beltrao said in a statement.
While AlphaFold helped Beltrao study the sequences between proteins in various species over time, the technique loses accuracy the further it regresses into the past.
6. Gauging the impact of rotavirus on gastroenteritis
Researchers at Baylor College of Medicine, Lawrence Berkeley Laboratory and Emory University School of Medicine are studying the differences in rotavirus groups A, B and C. While rotavirus group A and group C mainly infect young children, group B is primarily a problem for adults. In addition, rotavirus group B strains remain less well understood than the other strains. With AlphaFold, the researchers identified a new fold in rotavirus group B, potentially shedding light on why the strain tends to infect a different demographic than the other two strains.
7. Shedding light on Parkinson’s
An international research team has explored the use of AlphaFold to model the structure of stress-inducible phosphoprotein 1 (STIP1), exploring its role as a neuroprotective factor guarding against Parkinson’s disease. Currently, therapies that treat Parkinson’s tend to focus on symptomatic relief. No neuroprotective agents are approved to slow the rate of neurodegeneration associated with the disease. Researchers involved in the project were affiliated with Duke-NUS Medical School, Singapore General Hospital, A*STAR, Tan Tock Seng Hospital, National University of Singapore and Southern Medical University Guangdong.
Filed Under: Drug Discovery and Development, Infectious Disease





Tell Us What You Think!
You must be logged in to post a comment.